|
STORY CREDITS |
IITGN Researchers Develop An Open-Access Platform, Compiling Structures of Cancer-Linked Regulatory LncRNAs
Modern cancer research is advancing towards personalised medicine that addresses the unique molecular characteristics underlying an individual patient’s disease progression. To this end, scientists are exploring unorthodox and intricate regulatory layers that govern cancer cell behaviour. Among these, long non-coding RNAs, or lncRNAs, have attracted tremendous attention, considering their less-understood influence on cancer growth and progression.
To address the growing need for integrated studies on these regulatory RNAs, a team of researchers at IIT Gandhinagar (IITGN) has developed CanLncG4 (https://www.canlncg4.com), a comprehensive and user-friendly database that integrates structural and functional information on cancer-associated lncRNAs. The curated dataset available on CanLncG4 has been published in Scientific Data (Nature Portfolio) and offers a distinctive framework for studying the structure and behaviour of these RNA molecules.
“Long non-coding RNAs are an incredible class of RNA. While most RNAs mediate the translation of genetic code into proteins, lncRNAs exert protein-like functions such as guiding gene expression, helping repair damage, or influencing how cells grow,” explained Dr Bhaskar Datta, Professor at the Department of Chemistry and the principal investigator of the study. Dr Datta, who works jointly with the Biological Sciences and Engineering department at IITGN, added that when these molecules become dysregulated, which means their normal activity or levels are disturbed, they can be correlated with or cause tumour growth. Recent research has also shown that many lncRNAs fold into distinctive three-dimensional shapes that modulate their interactions with other molecules and, in turn, their regulatory functions.
One such shape is the G-quadruplex, or G4, a compact four-stranded structure formed when guanine, one of the four RNA building blocks, bonds in a particular pattern. These structures can act as tiny molecular switches that regulate how genes are expressed or silenced. When these switches malfunction, they can alter the behaviour of cancer cells. Understanding where and how these G4s form is, therefore, a critical step towards learning how lncRNAs influence cancer progression and drug resistance.
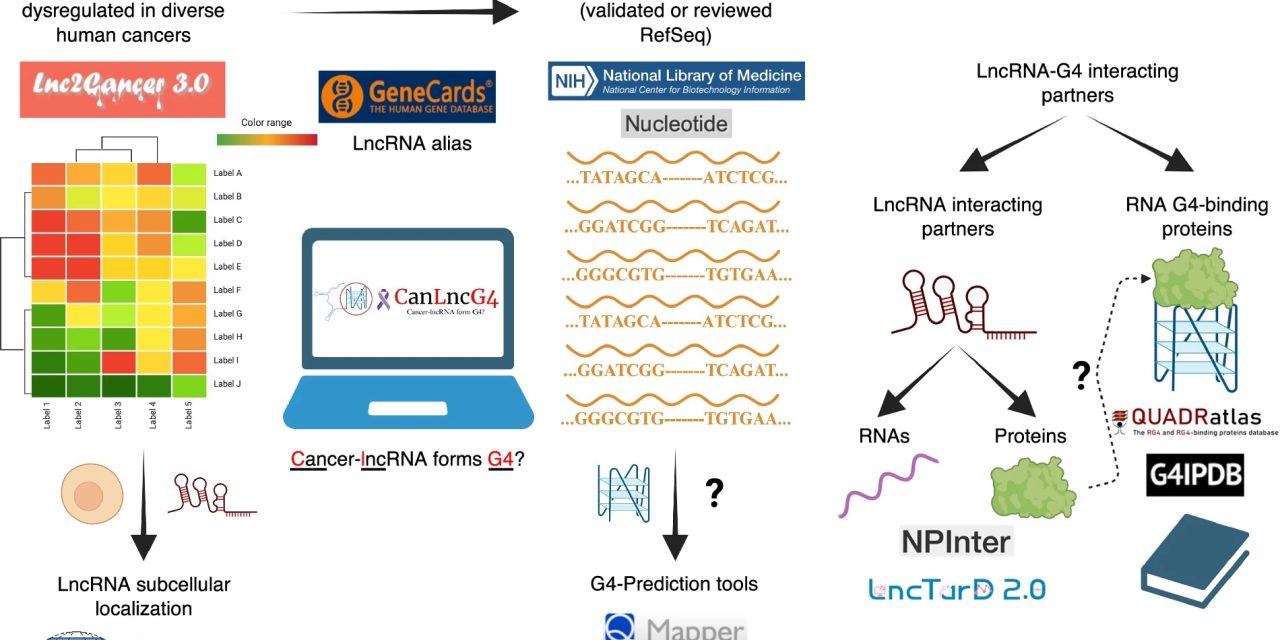
Existing datasets and informatics resources pertaining to dysregulated lncRNAs typically focus on individual features, such as RNA-cancer associations or RNA-protein interactions. This makes it difficult to study how these factors work together to regulate cellular functions, thus limiting their usability. CanLncG4 addresses this gap by integrating cancer-lncRNA associations, G4-forming potential, subcellular localisation, and molecular interactions in a single, user-friendly platform. It compiles experimentally validated associations between over 6,400 human lncRNAs and 15 cancer types, including breast, lung, liver, and prostate cancers. The dataset provides information on each RNA’s G4-forming potential and its location within the cell. “The open-access web interface of our dataset further enhances its utility by allowing researchers to search, visualise, and download data or identify new G4 predictions for RNA sequences of interest,” said Dr Shubham Sharma, the first author of the study. “Through its unique functional integration and accessibility, CanLncG4 serves not just as a data repository but as a powerful platform for hypothesis-driven investigations into cancer-associated lncRNAs and their structural regulatory roles,” he added.
By linking G4-rich lncRNAs to specific cancer types and their expression profiles, CanLncG4 offers valuable insights into how RNA structure influences tumour behaviour. This information can help identify potential biomarkers for early detection or prognosis and guide the search for new therapeutic targets based on RNA structure.
In addition to its biomedical applications, the dataset supports the growing field of precision medicine, where understanding the molecular architecture of individual tumours can inform tailored treatment strategies. “Our work builds on the growing recognition of non-coding RNAs in human health as highlighted by the 2024 Nobel Prize in Physiology or Medicine,” noted Dr Datta. “CanLncG4, as a structure-focused resource, highlights the role of RNA architecture in cancer and could accelerate both fundamental research and translational applications in the field.”
The other authors of the study include Muhammad Yusuf Hassan, Noman Hanif Barbhuiya, Ramolia Harshit Mansukhbhai, Dr Chinmayee Shukla, and Deepshikha Singh.